 Bugfixes in topo writelammpsdata (use resid instead of residue property, as basis for molecule ids, since this is what we read in) Bugfixes in topo readlammpsdata (can handle data files with class2 coefficients, doesn't writes topology information (bonds, angles, dihedrals, impropers) compatible with atom style) 
Bugfixes in topo guessatom (can handle element/type/name fields with unusual characters) - Bugfixes in topo writegmxtop (doesn't create charge groups with more than 32 atoms, better atom indexing) Bugfixes in topo readvarxyz (safer format checking, no spurious empty frames) Initial implementation for topo guessbonds (only works on "all" selections). Sanity check on atom coordinate data when reading lammps data files. This version contains important bugfixes in topo guessatom (one of which affects topo readvarxyz ), a few small enhancements, and a much improved and expanded documentation in addition to the tutorial on this homepage. The final version of TopoTools-v1.2 has been released with VMD version 1.9.1. 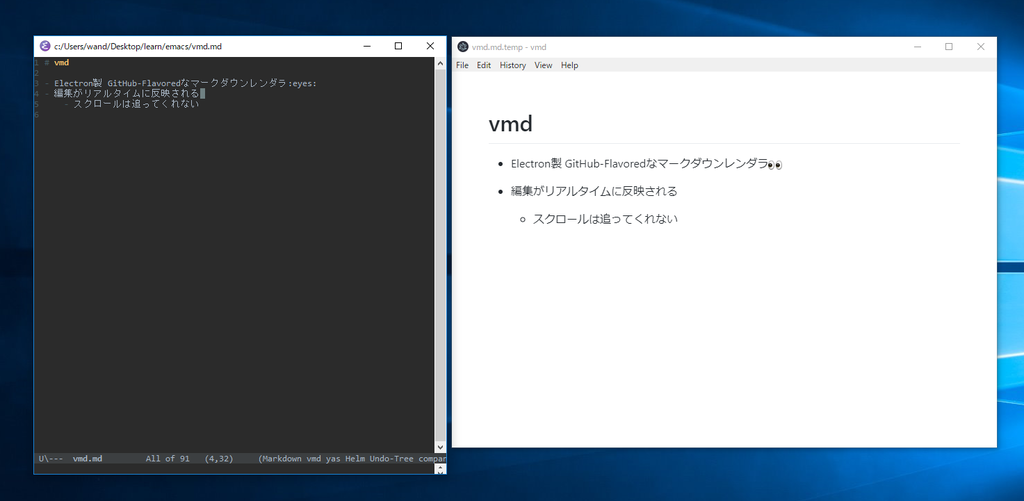
write commented out Coeff sections to data files to provide hints on which coefficients need to be set. Parse and compare - if present - against requested atom style on reading. encode the intended atom_style setting in data file header. only if the string CGCMM is in the first line of the data file, the lammps data file parser will look for those extensions. stricter checking for CGCMM extensions to the data file format. support for reading data files containing non-orthogonal cells. support for writing (fake!) gromacs topology files for use with gromacs analysis tools for non-gromacs trajectories. support for reading and writing of xmol/xyz style trajectories with a varying number of atoms element from mass, mass from element, element from name, radius from element). determination of atom properties based on other information (e.g. 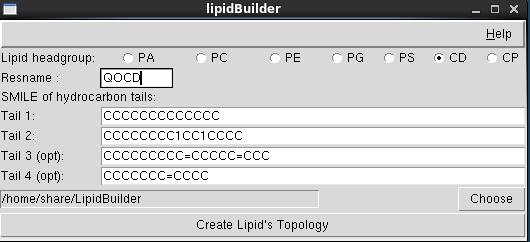
an optional tolerance keyword allows to specify the maximum difference from a flat structure. tested with peptide bonds, phenyls, histidines. searches for atoms bound to exactly three other atoms forming an almost flat construct. determination of angle and dihedral definitions from bonding information. Several bugfixes, especially for operating on selections and assigning atom properties. TopoTools-v1.1 has been released with VMD version 1.9. Key features: first public version, basic support for listing, typing, adding, and removing bonds, angles, dihedrals, and impropers, read/write support for LAMMPS data files, periodic system replication, merging of "VMD molecules" to a single molecule, merging of selections. TopoTools-v1.0 has been released together with VMD version 1.8.7. Please cite TopoTools as: Axel Kohlmeyer, (2017). #Install vmd how toLearn how to use TopoTools from the on-line tutorial. Please file bug reports and feature requests as GitHub issues. This should return the version number of the plugin that you have installed. To find out which version of topotools you have, please open the VMD text mode console and type: package require topotools. 
TopoTools consists of a generic middleware script layer that makes access to the topology related data stored in VMD more convenient than the existing very basic API, but it also has a number of high-level tools that allow reading and writing of topology file formats that cannot be parsed by the molfile plugins (as they need additional information not available to molfile plugins), parsing of parameter and residue database files for generation of complete input files for MD codes like LAMMPS and HOOMD-blue, and replicating or combining multiple systems (i.e. It is meant to be a complementary tool to psfgen, which is very much optimized for building topologies for biomolecules. TopoTools is a VMD plugin for manipulating topology information. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
